Simulation report
Technical information
General information
Name: Serotonin 5-ht1b receptor in complex with go protein
PDB id: 6G79
Activation state: Active
Submitted by: Nagarajan Vaidehi, Beckman Research Institute of the City of Hop
System setup
Solvent type: TIP3P
Membrane type: Homogeneous
Membrane composition: POPC
Ionic composition: Chloride (147 mM), Sodium ion (149 mM)
Number of molecules:
Water: 26041
POPC: 204
Sodium ion: 70
Chloride: 69
2-[5-[2-[4-(4-cyanophenyl)piperazin-1-yl]-2-oxidanylidene-ethoxy]-1~{h}-indol-3-yl]ethylazanium: 1
Guanine nucleotide-binding protein G(o) subunit alpha: 1
Total number of atoms: 119416
Simulation details
Software and version: GROMACS, 2016.4
Forcefield and version: CHARMM, CHARMM36
Time step : 2.0 fs
Delta : 0.2 ns
Replicates: 1
Accumulated simulation time: 1.797 µs
Additonal parameters: Available at "Simulation protocol & starting files"
References
Manbir Sandhu, Aaron Cho, Ning Ma, Elizaveta Mukhaleva, Yoon Namkung, Sangbae Lee, Soumadwip Ghosh, John H. Lee, David E. Gloriam, Stéphane A. Laporte, M. Madan Babu, Nagarajan Vaidehi. 2022. Dynamic spatiotemporal determinants modulate GPCR:G protein coupling selectivity and promiscuity. Nature Communications. doi: 10.1038/s41467-022-34055-5. (https://www.nature.com/articles/s41467-022-34055-5)
GPCRmd publication page: 1494
Simulation components
Ligands
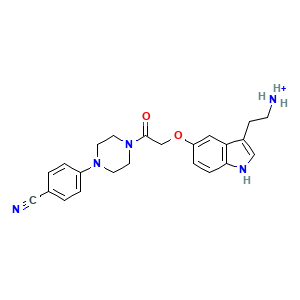
2-[5-[2-[4-(4-cyanophenyl)piperazin-1-yl]-2-oxidanylidene-ethoxy]-1~{h}-indol-3-yl]ethylazanium
(1 molecule)
Receptor
Mutations
Table of simulated mutations
| Residue ID | Residue seq. position | Generic num. | From | To |
|---|
Other molecules
2-[5-[2-[4-(4-cyanophenyl)piperazin-1-yl]-2-oxidanylidene-ethoxy]-1~{h}-indol-3-yl]ethylazanium (orthosteric lig.)
Specific State ID: 5321(1 molecule)